A new deep learning model, LucaProt, has detected over 251,000 new RNA virus species from global ecosystems, revealing unprecedented viral diversity in places like Antarctic sediment and extreme aquatic environments. Discover how this breakthrough could reshape our understanding of viral evolution.
Study: Using artificial intelligence to document the hidden RNA virosphere
In a recent study published in the journal Cell, researchers developed a deep learning model, “LucaProt,” a transformer-based AI model to detect highly divergent ribonucleic acid (RNA)-dependent RNA polymerase (RdRP) sequences in meta-transcriptomes from diverse ecosystems. They identified 180 RNA virus supergroups and 161,979 putative RNA virus species, showing that RNA viruses are widespread and present even in extreme environments.
Background
RNA viruses are widespread and infect a variety of species, yet their role in global ecosystems has only recently been recognized due to large-scale virus discovery efforts. These studies, primarily using RdRP sequences, have expanded the known virosphere by identifying thousands of new virus species. However, current tools often miss highly divergent RNA viruses, prompting the need for improved identification strategies.
Deep learning, particularly algorithms like convolutional neural networks (CNNs), recurrent neural networks (RNNs), and transformers, has revolutionized many areas of life sciences by offering more accurate and flexible approaches to identifying viruses. While CNNs and RNNs have been effective, they face limitations in processing long or complex sequences. Transformer architectures, which excel at capturing both short- and long-range relationships, present a promising alternative for discovering highly divergent RNA viruses.
Therefore, researchers in the present study developed a transformer-based artificial intelligence (AI) tool named LucaProt, which was rigorously benchmarked against several other virus discovery tools such as Diamond, HMMscan, HH-suite, and PalmScan. LucaProt achieved the highest recall rate (98.22%) and outperformed these methods in terms of recall and long-sequence processing. Additionally, LucaProt maintained a lower false positive rate compared to these tools to detect highly divergent RNA viruses from meta-transcriptomes to potentially reveal hidden viral diversity.
About the study
A total of 10,487 meta-transcriptomes, comprising 51 terabytes of sequencing data, were analyzed, of which 10,437 were obtained from the Sequence Read Archive of the National Center for Biotechnology Information database, covering diverse environments such as aquatic, soil, host-related, and extreme habitats.
Additionally, 50 datasets were generated from Antarctica and China, covering marine, freshwater, soil, and sediment samples. Sequencing and DNA/RNA extraction were performed. Sequence reads were assembled into contigs, and potential proteins were predicted using ORFfinder.
Two strategies were employed to identify potential viral RdRPs: LucaProt and ClstrSearch (a traditional approach that clusters proteins based on sequence similarity). The LucaProt model was trained on 235,413 samples, including 5,979 positive and 229,434 negative sequences, ensuring a comprehensive and well-validated dataset.
The results were compared to another method based on homologous protein clustering. A benchmarking test compared LucaProt with Diamond, HMMscan, HH-suite, and PalmScan tools. LucaProt outperformed these traditional tools, revealing significantly more new RNA viruses.
Reverse transcription polymerase chain reaction-based assays validated the presence of RNA organisms from viral supergroups. In addition, AlphaFold2 was utilized to predict the three-dimensional (3D) structures of viral RdRPs, with their structural similarities to known viral and eukaryotic polymerases thoroughly evaluated.
Results and discussion
LucaProt showed high accuracy (0.014% false positives) and specificity (1.72% false negatives). A total of 513,134 RNA viral contigs were identified using the two methods, representing 161,979 potential viral species (with over 90% RdRP identity) and 180 RNA viral supergroups, comparable to existing viral classifications by the International Committee on Taxonomy of Viruses.
Notably, LucaProt identified 70,458 putative unique viruses, including 60 previously unidentified supergroups, with the highest recall rate among all tested methods. Of these, 99.9% of viral contigs and 87.2% of supergroups were identified by both methods, while LucaProt identified an additional 444 contigs and 23 supergroups exclusively.
LucaProt achieved the highest recall rate of 98.22% among the tools. Other tools identified less than 42% of the new viruses exclusive to LucaProt. Notably, LucaProt recalled over 98% of RdRPs from other studies. Validation confirmed that the 180 new viral supergroups were RNA viruses based on RdRP motifs and sequence similarity.
Further analysis using AlphaFold2 revealed structural similarities between newly identified viral RdRPs and existing viral polymerases, enhancing confidence in the identification of novel RNA viruses.
The study also uncovered some of the most complex RNA virus genomes ever identified, including one genome that was 47.3 kilobases long, among the longest RNA viruses discovered to date. Most RNA virus genomes were around 2,131 nucleotides. Additional proteins were identified in new genomes, reinforcing their classification as RNA viruses.
The RNA virosphere expanded significantly, with a 55.9-fold increase in species compared to previous classifications. High phylogenetic diversity was found in newly discovered supergroups, indicating potential for more divergent RNA viruses.
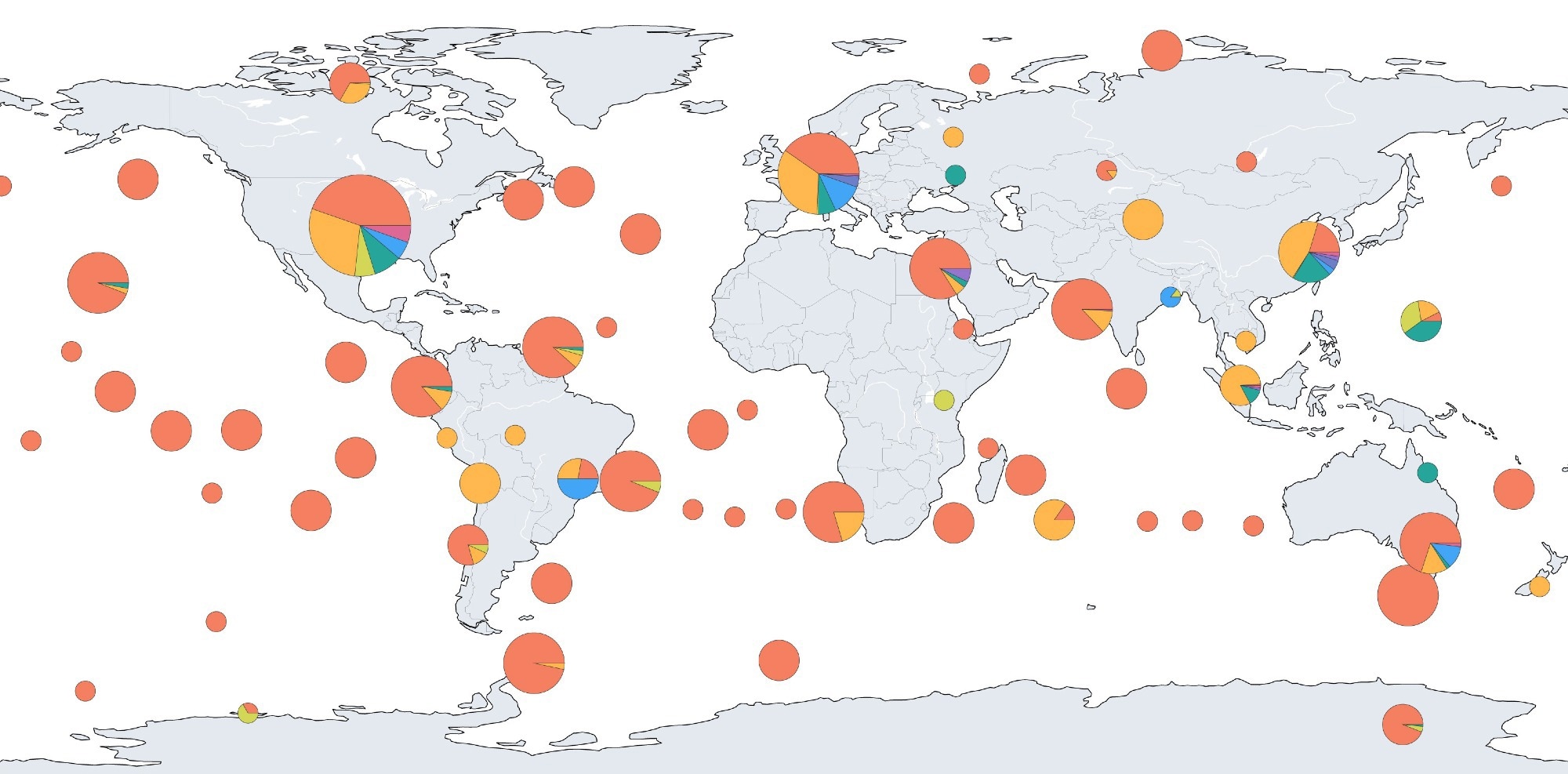
Widespread virus presence was revealed across 32 ecosystem subtypes and 1,612 locations, with 33.3% of groups identified by LucaProt being previously unreported. Alpha diversity, a measure of species diversity within an ecosystem, was highest in environments like leaf litter, while viral abundance peaked in Antarctic sediment and marine environments.
Many new viral supergroups were predominantly aquatic or sediment-based, with a few associated with specific host ecosystems. However, systemic biases in data generation may affect comparisons across ecosystems. The study’s limitations include challenges in classifying highly divergent viruses, lack of matching DNA data for some virus groups, and the identification of only partial viral genomes focused on RdRP segments.
Conclusion
This study enhances our understanding of the RNA virosphere by identifying over 251,000 new viral species and 180 novel supergroups using deep learning and large-scale meta-transcriptomic analysis. These findings highlight the vast genetic diversity of viruses in environmental samples, emphasizing the importance of ongoing research in ecology and public health related to viral pathogens and ecosystem dynamics.
Source link : News-Medica